SLICSel Help
Overview of the program
SLICSel is a program for designing specific oligonucleotide probes for microbial detection and identification. To obtain the maximal specificity of designed oligonucleotides, SLICSel uses the Nearest-Neighbor thermodynamics-based approach for probe design.
SLICSel will try to design reverse complement probes for capturing the target sequence(s) so that designed probes would not cross-hybridize with the control sequences.
Target sequence(s): the sequence(s) one wishes to capture with the oligonucleotide probes designed with the SLICSel program. In case of several target sequences, SLICSel will try to design a common capture probe for all those sequences.
Control sequence(s): the sequence(s) which should not hybridize (in specified experimental conditions) with the probes designed by the SLICSel program.
Probe melting temperature interval: a range of temperature into which SLICSel will try to design unique probes. The melting temperature is calculated by using the following formula:
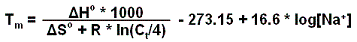
Probe annealing temperature: temperature calculated from the melting temperature interval using the formula below:
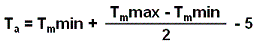
Evaluated annealing temperature will be used to correct the thermodynamic calculations.
Minimal binding energy (ΔG) difference between target and control sequences: the minimal binding energy which should be between the correct binding with the target sequence and the strongest (non-specific) binding within the control sequences.
Maximal binding energy (ΔG) difference between target sequences: the maximal binding energy difference inside the target group for one probe that should capture all target sequences.
Thermodynamics parameters for probe design: target sequence dependent parameters for oligonucleotide probe design. If one would like to capture RNA molecules it is recommended to use the DNA-RNA Nearest-Neighbor parameters for probe design. In case of DNA target molecules, it is recommended to use the DNA-DNA parameters for probe design.
For further information please contact:
